Dr Lisa Connor is head of the Vaccine Evaluation team for the Vaccine Alliance Aotearoa New Zealand - Ohu Kaupare Huaketo. She completed her PhD at the Malaghan Institute with Associate Professor Jo Kirman and Postdoctoral Fellowship at the Trudeau Institute, NY, USA, focused on vaccination against respiratory diseases tuberculosis and influenza.
After returning to New Zealand in 2012, her research interests turned to dendritic cells, a major target for vaccine development and investigated dendritic cell instruction of immune cell fate with Professor Franca Ronchese at the Malaghan Institute. In 2017 she was awarded a Sir Charles Hercus fellowship from the Health Research Council of New Zealand to examine the impact of the tissue microenvironment on dendritic cell function. In 2018, Lisa established her research lab in the School of Biological Sciences at Te Herenga Waka—Victoria University of Wellington.
Research interests
"I am a cellular immunologist with an interest in the host-pathogen interface. Our immune systems play a remarkable role in the defence against microscopic villains, and can orchestrate a faster superior response upon re-exposure, which is the basis of vaccination. Our research focus is to investigate novel vaccines that promote potent humoral and cellular immunity in the respiratory tract – a site where many pathogens try to invade."
Research projects
- Identify effective vaccine candidate for SARS-CoV2 virus
- Target lung resident lymphocyte populations to develop mucosal vaccines for respiratory viruses (i.e. influenza)
- Investigate B cell responses, antibody production and T follicular helper response to vaccines that harness tissue resident innate-like lymphocytes
- Identify the impact of the skin and lung microenvironment on dendritic cell function and heterogeneity of adaptive immunity
- Identify key molecules involved in dendritic cell: T-cell communication
Harnessing innate like T-cells as mucosal adjuvants against respiratory pathogens
Contributors – Ian Hermans, Gavin Painter, Theresa Pankhurst, Kaitlin Buick, Olga Palmer, Joshua Lange, Benji Compton, Sarah Draper
Over 90% of pathogens invade the human host through mucosal routes. As a result, our immune system has created a specialised mucosal immune response, armoured with cells and immunoglobulins (Ig) designed to defend mucosal surfaces. Such defences include production of secretory IgA (SIgA), which block pathogen invasion, and the presence of long-lived tissue resident memory T cells. Despite the specialised role of mucosal immunity, the majority of human vaccines are administered parenterally and fail to elicit mucosal immune responses.
New vaccine designs are moving away from whole cell-based approaches. Instead, the less immunogenic subunit vaccines that require an “adjuvant” - compounds added to a vaccine that specifically enhance induction of adaptive immune responses, are favoured. This poses a significant barrier for mucosal vaccines as 1) many adjuvant formulations are not appropriate for mucosal delivery; 2) mucosal surfaces act as a physical barrier that can prevent effective access to the target immune cells. We propose that we can overcome the current barriers for mucosal vaccine development by designing adjuvants that target innate-like T cells located in the mucosal tissue. In collaboration with Prof Ian Hermans, MIMR, Prof Gavin Painter, Ferrier Research Institute and Avalia Immunotherapies we are designing and testing novel mucosal vaccines for influenza virus.
Deciphering the unspoken language between immune cells
Contributors – Davide Comoletti, Cynthia Morgan, Liam Turk
Communication between dendritic cells (DC) and T-cells is essential for activating the adaptive immune response. Information passed on from the DC to the T-cell is typically mediated by receptor-ligand interactions. As a result, proteins expressed at the DC T-cell interface provide an attractive target for therapeutic intervention. DCs are well-known for their ability to sense their environment, and in turn instruct T cells to respond most appropriately to a given situation. However, in the case of allergy, T-cells (following instruction by DCs) inappropriately target harmless substances and differentiate into T helper (Th) 2 cells. This project explores receptor ligand interactions that occur between DCs and T-cells. In collaboration with VUW molecular protein biologist, Dr Davide Comoletti, we are developing a high-throughput screen that has the capacity to go beyond a single interacting pair and feasibly screen >80,000 interactions. The ultimate goal of this screen is to identify novel protein-protein interactions between DC and T cells and determine key molecules involved in Th2 development.
PhD (Otago) BMedSc(Hons) (VUW)
Related news

Malaghan scientists appointed in leadership roles in New Zealand's RNA Platform
31 May 2024
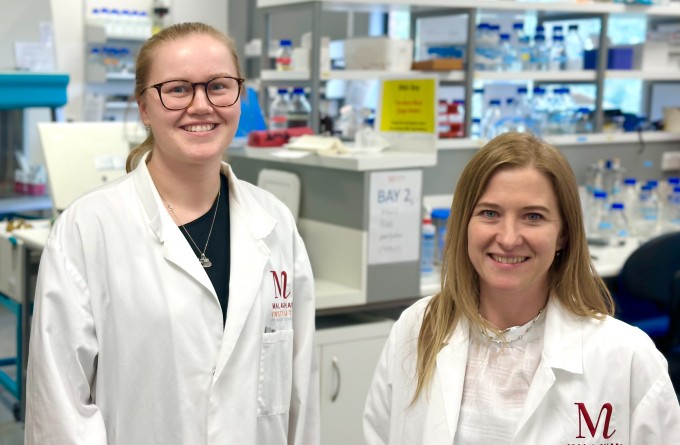

Stopping infections at the door – designing vaccines that target immune cells in the mucosa
19 April 2023
Kiwi-made Covid-19 booster vaccine offers 100% protection in preclinical study
3 March 2023

Homegrown COVID-19 booster vaccine: building New Zealand’s biomedical capability
23 May 2022

Discovery points to the skin as ‘ground zero’ for allergic disease
19 November 2021

Vaccine alliance formed as collaborators establish national COVID-19 vaccine screening and development programme
27 August 2020
Publications
2024
Pankhurst TE, Montgomerie I, Marshall A, Draper SL, Bilbrough T, Button KR, Palmer OR, Hermans IF, Painter GF, Connor LM, Compton BJ (2024). A Glycolipid-Peptide-Hapten Tricomponent Conjugate Vaccine Generates Durable Antihapten Antibody Responses in Mice. ACS Chem Biol. 19(6):1366-1375
2023
Pankhurst TE, Buick KH, Lange JL, Marshall AJ, Button KR, Palmer OR, Farrand KJ, Montgomerie I, Bird TW, Mason NC, Kuang J, Compton BJ, Comoletti D, Salio M, Cerundolo V, Quiñones-Mateu ME, Painter GF, Hermans IF, Connor LM (2023). MAIT cells activate dendritic cells to promote TFH cell differentiation and induce humoral immunity. Cell Rep. 42(4):112310
Montgomerie I, Bird TW, Palmer OR, Mason NC, Pankhurst TE, Lawley B, Hernández LC, Harfoot R, Authier-Hall A, Anderson DE, Hilligan KL, Buick KH, Mbenza NM, Mittelstädt G, Maxwell S, Sinha S, Kuang J, Subbarao K, Parker EJ, Sher A, Hermans IF, Ussher JE, Quiñones-Mateu ME, Comoletti D, Connor LM; On behalf theVAANZ Group (2023). Incorporation of SARS-CoV-2 spike NTD to RBD Protein Vaccine Improves Immunity Against Viral Variants. iScience. 26(4):106256
2021
Mayer JU, Hilligan KL, Chandler JS, Eccles DA, Old SI, Domingues RG, Yang J, Webb GR, Munoz-Erazo L, Hyde EJ, Wakelin KA, Tang SC, Chappell SC, von Daake S, Brombacher F, Mackay CR, Sher A, Tussiwand R, Connor LM, Gallego-Ortega D, Jankovic D, Le Gros G, Hepworth MR, Lamiable O, Ronchese F (2021). Homeostatic IL-13 in healthy skin directs dendritic cell differentiation to promote TH2 and inhibit TH17 cell polarization. Nat Immunol. 2021 Nov 18
